 |
| Illustration: Andrea Dingeldein |
Microbial DNA reveals invisible societies that control the world’s oceans.
Lindzi Wessel wades into the science. Illustrated by Andrea Dingeldein.
At 8 a.m. every Wednesday, UC Santa Cruz lab technician Thea Fredrickson goes to the Santa Cruz Municipal Wharf and dons her poop-jacket. The black windbreaker lets her stretch her arms far out over the wharf’s railing without getting bird droppings and fish guts on her clothes. From 20 feet above Monterey Bay, she lowers a white five-gallon bucket into the ocean and reels it back up with practiced hands. A few inches of water slosh in the pail.
Her partner, undergraduate student Sanjin Mehic, adds two more splashes of seawater from a small gray canister that he lowered below the surface. The canister went down twice, collecting water from about 15 and 30 feet deep. The labmates are capturing representative samples of three layers of the upper ocean. They will bring the clear, cool water to several research labs on campus, where scientists care not about the water itself, but the bustling and diverse microscopic societies it contains.
 |
Invisible to our eyes, these communities have enormous power in their sheer numbers, says UCSC ocean scientist Jonathan Zehr, a recipient of the bay’s water. Each Gatorade-size bottle can contain about half a billion microbes and as many as 40 million unique DNA sequences. This represents a minuscule fraction of the marine microbiome: myriad cells that breathe life into the world’s oceans. Each microbe is hundreds of times smaller than a human cell, but together these plants, viruses, bacteria and other life forms make up more than 90 percent of the living matter in the sea.
Bacteria are the most abundant, says Zehr. One teaspoon of seawater can contain at least half a million bacterial cells. “If you evaporated the ocean,” he says, “the biggest piles wouldn’t be the whales. They would be the piles of bacteria.”
The marine microbiome remains mysterious; most of its cells are too small to isolate and grow in the lab. But advances in genetic technology have provided a new way to explore what’s out there. Robotic underwater labs and ocean voyages to collect DNA are helping us grasp the diversity of the society that keeps life on Earth afloat. In an improbable quest to track communities of microbes across the roughly 300 million cubic miles of water that make up the planet’s oceans, researchers like Zehr are studying how many species of these microbes exist, how they interact, and how they influence the ocean’s health. Along the way they have made some startling discoveries, upending our basic understanding about what it means to be alive in the sea.
“There’s a whole lot of life out there,” Zehr says. “We don’t even know what it is.”
Sea cells
When Zehr began studying aquatic invertebrates as a graduate student in the early 1980s, scientists were just starting to learn the roles played by microorganisms in watery realms. New developments that allowed researchers to track energy and nutrient cycles showed that microbial communities, not invertebrates or other animals, drove the metabolism of aquatic ecosystems. “I realized that what was going on with the microbes was so much more important,” says Zehr. “So I changed my topic and started becoming a microbial ecologist.”
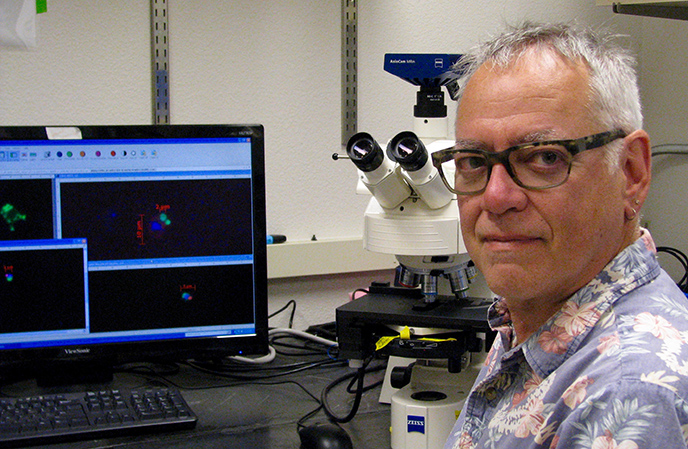 |
| Photo: Lindzi Wessel |
| UC Santa Cruz ocean scientist Jonathan Zehr in his lab. |
|
As his career developed, Zehr turned to what may be the most influential microbiome on the planet: the ocean’s. A continuously shifting soup covering almost 75 percent of the planet, the ocean provides a home for enough microbes to produce an estimated half of all oxygen on Earth. They control the balance of gases in our atmosphere and form the foundation of the sea’s food web.
Like farmers try to understand how the invisible communities in soil will influence their crop yields, Zehr wants to know how the microscopic marine societies influence the ocean’s fertility. Farmers know that rotating their harvests with crops like peanuts, soybeans, and other legumes will fertilize their fields. This ability to rejuvenate soil comes from bacteria living in the plants’ roots. The bacteria pull nitrogen from the air and convert it into a form that plants need to survive. Recently, researchers found that legume roots do not have a monopoly on these special organisms, called “nitrogen fixers.” Their microbial kin populate the ocean.
Nitrogen fixation, Zehr says, is essential for marine life to thrive. But to study these unique microbes, Zehr and his colleagues first must find them. The search compels them to take to the sea.
Zehr spent much of his early career on ships. Now, he regularly sends his team to collect microbes from the open ocean. One target of these voyages is the North Pacific Gyre. Spanning most of the Pacific Ocean, this vast whorl of currents defines the largest continuous ecosystem on Earth and hosts a broad diversity of microbes.
In particular, the waters near the Hawaiian Islands are a common destination for researchers to track aquatic microorganisms. Just as tropical landscapes of Hawaii seem nothing like the redwood forests near Santa Cruz, the Hawaiian seas contain habitats much different than the one Fredrickson samples from the wharf each Wednesday.
During cruises near Hawaii, patches of color often pass under the ships—colonies of microbes so large that satellites can see some clusters from space. Massive stretches of algae with nitrogen fixers tucked inside their cells can tint the water orange, red, or green. In some cases, satellites detect clusters of Trichodesmium, a cyanobacteria nicknamed “sea sawdust” because of its propensity to coat the surface of the ocean with streaks of clumping colonies. These chains of cells sink or rise through the water to chase sunlight and nutrients by shifting sacks of air inside each cell.
But Zehr also hunts for elusive and poorly understood cousins to Trichodesmium that don’t group together and remain too small to be seen. If a billion of these loner cells were spread evenly in a liter of seawater, each would be separated from its nearest neighbor by 200 to 300 body lengths. Scaled up to the size of a person floating in an inner tube, each microbe would find itself more than five Olympic swimming pool lengths away from the next tuber. To find the cells, Zehr paradoxically has turned to something even smaller: the genes in their DNA.
Zehr and his colleagues use genes like animal trackers use footprints. They serve as clues that tell researchers what kinds of organisms have been in an area and what those organisms have been doing. The scientists’ high-tech sampling bucket is a large ring of metal canisters that a sturdy crane lowers below the waves. This ring is fitted with sensors to keep track of the water’s conductivity, temperature, and depth—hence its name of “CTD rosette.” The rosette plunges deeply to collect samples at different levels in the water. In the lab, researchers scour these samples for interesting genes.
Scientists have long been able to detect specific genes by creating probes that bind to them. Like two strands of a zipper combining to form a unified chain, the probe sticks when it finds its partner. Researchers engineer the zipper strand to glow if the genes they seek are there. If they aren’t, the probes don’t stick. The glowing probes point scientists to important genes, like those that allow microbes to convert sunlight into energy or nitrogen into food for aquatic plants.
On a research cruise in 1998, Zehr plucked a strange nitrogen-fixing gene out of the sea. At the time, he didn’t have the tools to detect the organisms the gene belonged to. But the genetic imprint told him that an unidentified species of cyanobacteria—formerly called “blue-green algae”—lurked in the water.
Zehr started to see these luminescent probes pointing to his mysterious nitrogen-fixing gene in water samples from around the globe. Not knowing exactly what it was, marine microbial biologists endowed it with a temporary name: “UCYN-A,” for unicellular cyanobacterium nitrogen-fixing-group A.
For years it seemed obvious that the key nitrogen fixers were the ones that could be seen; massive clusters held huge numbers of them. But as the genetic footprint of UCYN-A popped up on more and more research voyages, Zehr began to suspect that this organism, though invisible, was making a substantial contribution to life in the oceans.
But Zehr couldn’t understand his quarry until major advances in genetics led to tools that analyzed the individual components of DNA. Now, instead of looking for the long DNA sequences of a few special genes, researchers can read every single genetic unit of every single sequence. Just as identifying each letter allows a new reader to sound out unfamiliar words, this new technology meant scientists could read every gene they pulled out of the ocean.
|
| Graphic: Lindzi Wessel |
| Research cruises have detected nitrogen-fixing microbes in the global ocean along a growing number of sampling pathways (green dots). Zoom in to see cruise tracks in more detail. |
An ocean of data
Within just a few years, genetic sequencing methods flooded the field of microbial ecology with more data than scientists had power to process. “I started my Ph.D. almost eight or nine years ago,” says Hanna Farnelid, a postdoctoral researcher studying microbes in Zehr’s lab. “Back then we were still studying hundreds of sequences. Now, it’s millions and billions.”
Insights from this deluge of data gave Zehr new clues about UCYN-A, but they also deepened the decade-long mystery. In a stark surprise, the organism’s whole genome was missing most of the fundamental genes that support life. There was only one explanation: the thriving microbe has a mystery partner. “We don’t know who it’s working with,” Zehr says, “but it’s working with someone.”
So Zehr went on the hunt again, this time to expose UCYN-A’s enigmatic comrade. Preliminary genetic data suggests it is a new species of microscopic algae. But even as Zehr closes in on this second genome, he and his colleagues expect more surprises to surface.
“We’re making exciting discoveries like this every year, because there are still large unknowns about how things work together [in the sea] and how these organisms live,” says Farnelid.
One thing is clear: marine microbes don’t play by the straightforward rules scientists once ascribed to them. The community is full of interaction and drama, says Mary Ann Moran, a microbial ecologist at the University of Georgia. These simple cells, many just specks under a microscope, are constantly developing innovative strategies to survive. Her favorite example is a bacteria that “farms” the leaky, microscopic plant cells it turns to for food. The bacteria releases hormones that cause the plants to grow, allowing it to consume more of the organic material that seeps out of the cells. “It’s almost diabolic,” Moran says.
The expansive genetic passages within marine life forms are revealing many such untold stories. “People literally just go in and pull out one cell—one single cell—and get a genome sequence out of it,” says Moran. “We’re not stuck anymore. If it won’t grow for us in the lab, it doesn’t mean we can’t learn an amazing amount about its biological and ecological roles.”
Sensor network
Researchers on ships cannot sample quickly enough or with great enough range to capture fine details from the sea’s continuously shifting microbial world. To cover more territory, Zehr’s lab is taking on some unusual personnel: robots.
“There really is a need for a persistent presence, and it’s not sustainable with boats and people,” says marine microbiologist Chris Scholin. Scholin and his team at the Monterey Bay Aquarium Research Institute are providing marine ecologists like Zehr with autonomous underwater robots. The machines perform laboratory-quality tests directly in the ocean. Deep beneath the waves, the robots collect samples, burst cells open with heat and chemicals, and test the genes that spill out.
|
Lindzi Wessel visits the Monterey Bay Aquarium Research Institute to talk with an engineer about robotic probes designed to study microbial life in the ocean. Click on image to play. |
MBARI engineers are working on a new line of small, mobile labs to track microbial communities. Small enough to live in the nose of an unmanned glider, the labs can drift along with clusters of microbes like ethnographers quietly shadowing the societies they study. The ten-foot-long torpedoes will let researchers watch these dynamic systems of organisms change from the microbes’ frame of reference, rather than grabbing a small piece of a sample drifting past.
Scholin and Zehr hope genetic tools, coupled with new sampling techniques, will help scientists make predictions about the future of the oceans. Tracking changes in microbial communities, Scholin says, should yield clues about the health of marine ecosystems. The goal is to distill the behavior of millions of genes into a handful of important indicators that will help scientists understand how ocean waters are mixing, which nutrients are available, and how the marine food web might shift.
“[Microbes] are tuned exquisitely for all sorts of things that aren’t easy to measure,” Scholin says. “If we could ever learn how to read them, they could be the greatest sensor network out there.”
Microbial legacy
But even as ocean biologists delight at the growing access they have to marine secrets, they worry. The seas are changing as they work. Earth is warming, and the ocean is heating up. Excess carbon dioxide from global emissions enters the water as well, making seawater more acidic. Increasingly corrosive water poses a new challenge to many organisms. For many researchers, these changes have added urgency to their quest to understand ocean ecosystems and how shifts in microbial communities will affect them. “Nobody really knows what is going to happen,” says Scholin.
The ways microbes respond to changing conditions may have impacts up and down the marine food chain, he says. Damage to coral reefs and increased diseases in marine organisms may both be catalyzed by changes in the structure and function of microbial communities. “If you step back and add it up, suddenly you realize those things have a lot of power over us,” says Scholin. “There are scenarios where people project that given enough time and with the right conditions, the microbes could transform huge swaths of the ocean.”
It has happened before. Two billion years ago, microbial communities altered the planet when they began producing oxygen—setting the stage for multicellular life to arise. No matter how marine ecosystems have evolved since then, these ancient beings have been in the driver’s seat. Zehr doesn’t expect this to change. “They control the Earth,” he says. “They'll be here long after we're gone.”
© 2016 Lindzi Wessel / UC Santa Cruz Science Communication Program
Top
Biographies

Lindzi Wessel
B.S. (psychology) University of California, Davis
M.S. (neuroscience) University of California, Davis
Internship: STAT / Boston Globe
At the dinner table my family talks politics. A lot. I was more interested in how the brain forms, so I turned on the microscope instead of the news. (I also scoured the cognition literature in vain for strategies to tune out parental guilt.) But while studying how neurons grow using adult human stem cells, I started to explore the fraught history of stem cell research. I couldn’t believe how much our political and legal systems influence scientific advancements—and how science policies arise with so little public input.
Once I began to investigate the societal relevance of research around me, I couldn’t stop. Now during family visits, I’m talking about science and democracy. A lot. As a science writer, I hope to spark many more of these conversations, one dinner table at a time.
Lindzi Wessel’s website
. . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . .

Andrea Dingeldein
B.S. (marine biology) and B.A. (studio art) University of North Carolina Wilmington
M.S. (marine biology) University of North Carolina Wilmington
Internship: Friday Harbor Laboratories, University of Washington
Equipped with a strong background and several degrees in marine biology, Andrea mostly focuses her science illustration work on subjects from the marine world. Her experiences as a diver and kayaker give her a unique perspective and enable her to represent the underwater environment and creatures with great accuracy. She enjoys the play of light and shadow and uses them to create drama and highlight areas of interest in her illustrations. Andrea works with both traditional and digital media and enjoys partnering with scientists to create unique infographics that disseminate scientific research to a broader audience.
Andrea Dingeldein’s website
Top |
